Engineered systems for studying cell fate and function
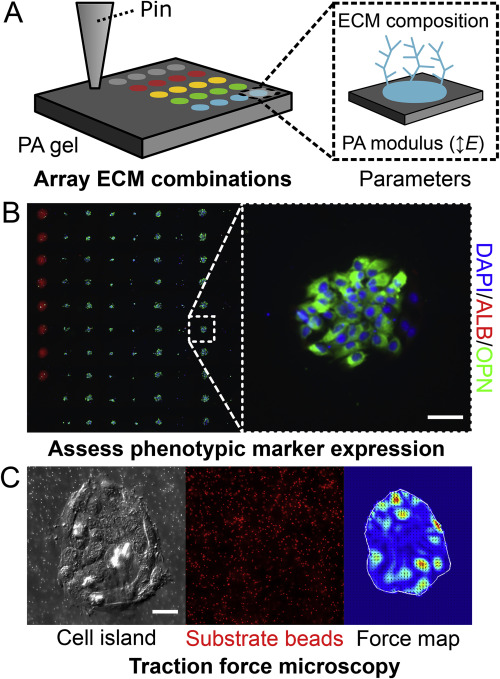
Cells are exposed to a variety of regulatory cues in their in situ microenvironment. These microenvironmental cues include cell–cell, cell–soluble factor, and cell–matrix interactions as well as more fundamental physicochemical considerations such as pH, mechanical forces, and electrical potential. Efforts utilizing engineered systems have been focused on recapitulating these cues to attain fine control of cellular processes such as fate specification. In order to systematically investigate these cellular processes, my work in this area has focused on designing engineered systems capable of high-throughput presentation of microenvironmental signals, such as extracellular matrix proteins or signaling ligands. I have been particularly interested in enabling investigations of combinatorial regulation of cell fate through the integration of additional readout modalities and analytical approaches with these engineered systems.
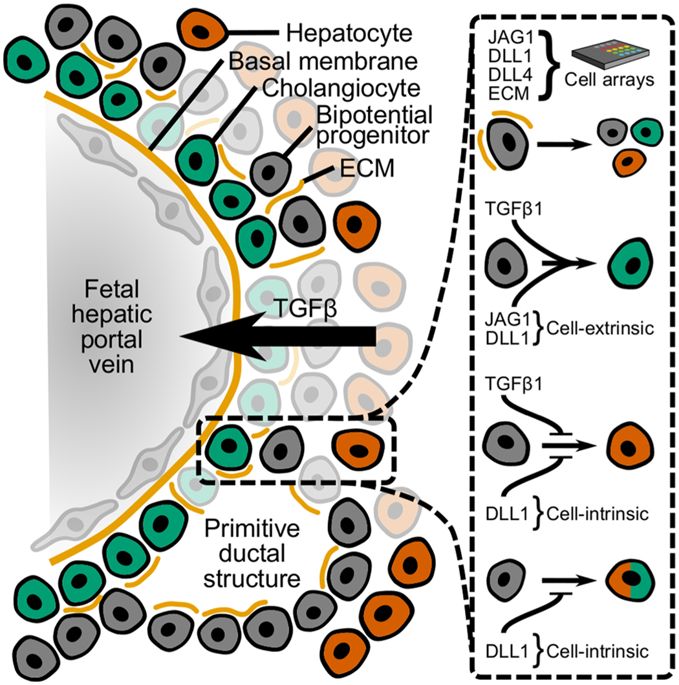
Liver development and bile duct morphogenesis